We build quantitative models informed by molecular and epidemiological data to guide the public health response to emerging and endemic pathogenic threats.
The Martin Group @ JHU BSPH is a computational biology research group in the Department of Epidemiology at the Johns Hopkins Bloomberg School of Public Health. Broadly, our work aims to reduce the global burden of infectious diseases. More specifically, we use pathogen molecular (particularly genomic) data and quantitative mathematical models to better understand the biological, environmental, and sociological drivers of pathogen population dynamics to inform the public health response to endemic and emerging infectious threats. We are a part of the Infectious Disease Dynamics group at JHU and collaborate broadly throughout the university and beyond.
Most of our work falls into one of the following themes:
Within-host pathogen evolution
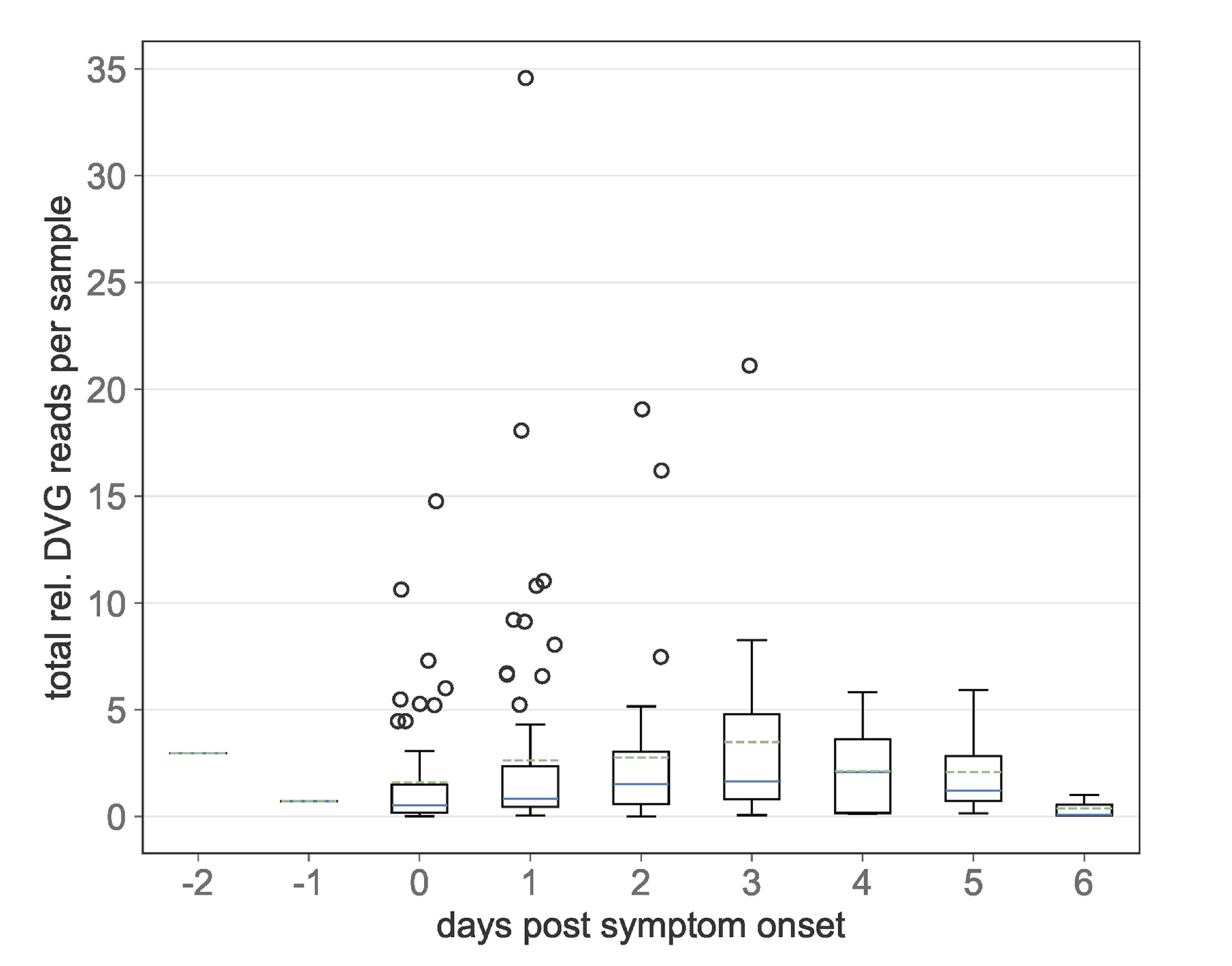
All pathogen genetic diversity observed at the host population scale must first be generated within individual hosts. We use pathogen deep-sequence data and evolutionary models to better understand the emergence and maintenance of within-host pathogen genetic diversity and implications for long-term pathogen evolution.
Cross-scales pathogen diversity
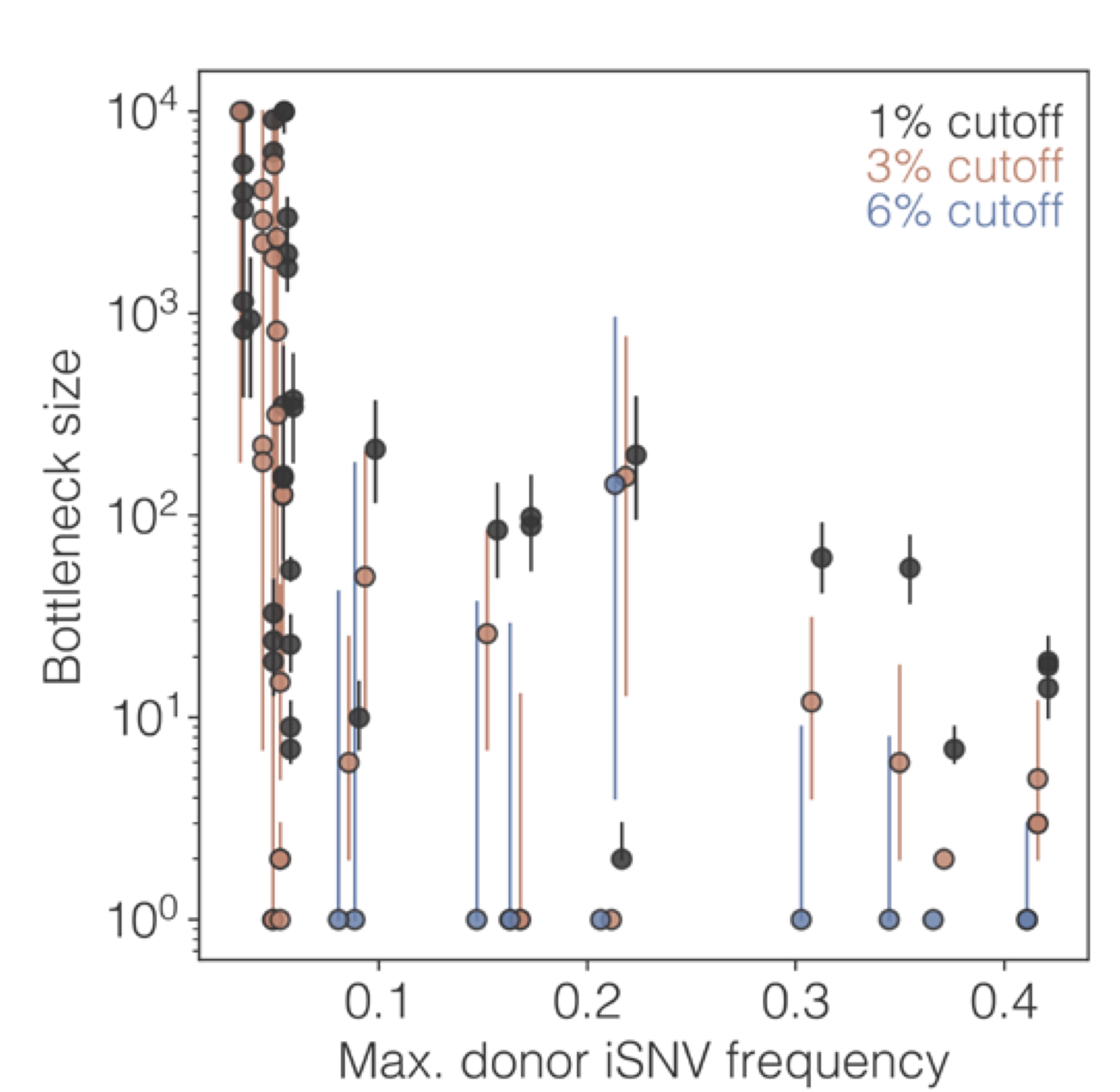
Pathogen genetic diversity generated within-hosts must transmit in order to persist at the population scale. We fit models to pathogen deep-sequence and well-resolved epidemiological data to better understand the biological dynamics governing these cross-scales dynamics.
- Relevant publications:
- SARS-CoV-2 bottleneck
Population-scale pathogen dynamics and population genetics
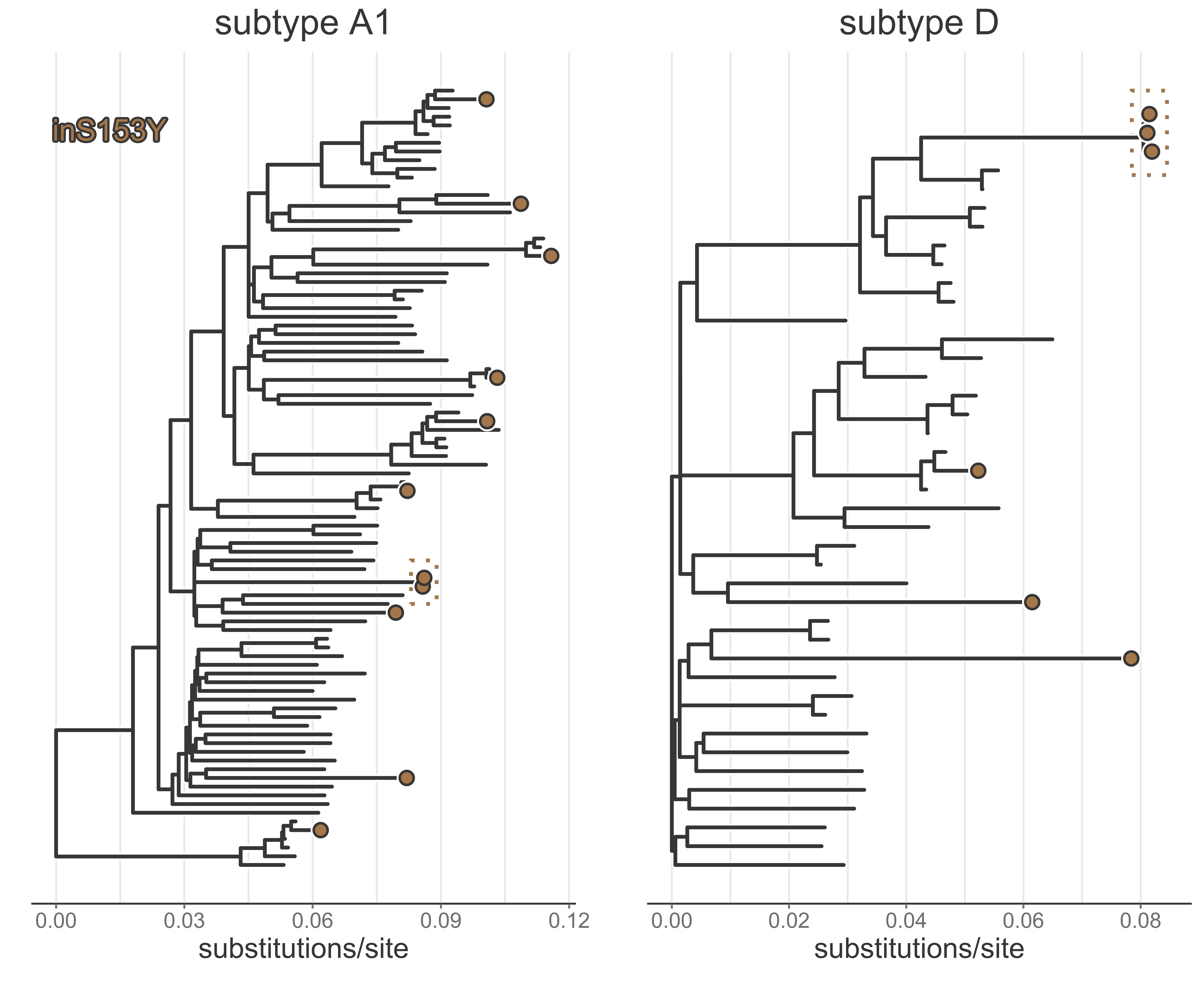
As epidemiologists we are ultimately interested in pathogen dynamics at the host population scale. To this end we pair pathogen sequence data with detailed metadata and models of the underlying dynamics to understand the evolutionary (e.g. fitness evolution, treatment resistance) and exogeneous (e.g. climate, host behavior) factors shaping pathogen population dynamics. Much of this work uses phylodynamic approaches but we also have ongoing work in developing tree-free methods.
Ecological interactions in multi-pathogen systems

Many pathogens share ecological niches e.g. sexually transmitted pathogens, respiratory pathogens, mosquito transmitted pathogens. However, little is known about how the direct and indirect interactions between co-circulating pathogens and the impact of interventions targeting one pathogen on the dynamics of their ecological neighbors. In this emerging research theme, we are combining sequence data from co-circulating pathogens with ecological models to better understand these interactions.
